葉綠體DNA
葉綠體DNA(Chloroplast DNA,cpDNA)為真核生物細胞中葉綠體內的DNA。葉綠體等质粒体中具有一獨立於細胞核的基因組,且含有核糖體可轉譯合成自身的蛋白質[1][2],為一半自主的胞器,葉綠體DNA最早於1959年透過生化實驗發現[3],並於1962年透過電子顯微鏡實際觀測確認[4]。1986年菸草與地錢的葉綠體DNA被定序發表,為最早完成定序的葉綠體基因組[5][6],目前已有大量陆生植物與藻類的质粒体被定序發表。
基因组結構
编辑葉綠體DNA為環狀,長度一般介於120至170kb之間[7][8][9],分子量為8000萬至1.3億Da[10],多數植物的葉綠體包含約120個基因[11][12],大多編碼光合作用所需的蛋白(包括RuBisCO的大次單元)與基因表現所需的蛋白[13],如4種rRNA、約30種tRNA、21種核糖體蛋白以及葉綠體RNA聚合酶的4個次單元[13]。絕大多數生物的葉綠體DNA均不分段,雙鞭毛蟲門的藻類則為罕見例外,此類生物的葉綠體DNA包括約40段質體,每個質體長2至10kb,包含1至3個基因,也有些質體不編碼任何基因[14]。
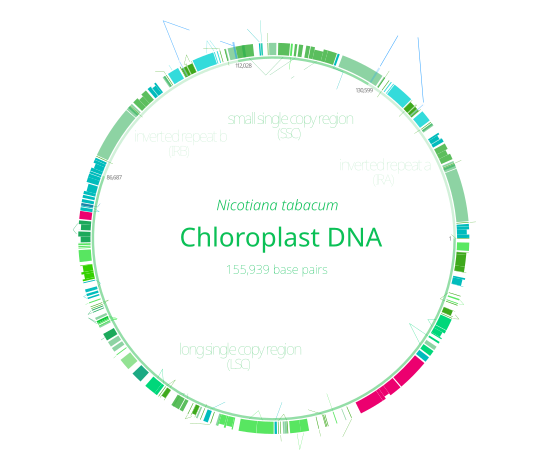
多數葉綠體DNA包括兩段反向重複序列(IRa與IRb),將葉綠體DNA分成大單拷貝區(LSC)與小單拷貝區(SSC)兩區域[9]。反向重複序列長4至25kb,其中植物的一般在20至25kb之間[15],IRa與IRb的序列一般會因協同演化而非常近似[14]。植物葉綠體DNA的反向重複序列演化上相當保守[9][15],藍菌基因組與灰藻和紅藻的葉綠體基因組中也有與其同源的序列,顯示此序列在演化上的起源很早,在葉綠體出現前即已存在[14]。有些葉綠體(豌豆與數種紅藻[14])的基因組丟失了反向重複序列[15][16],紫菜屬紅藻葉綠體則有一個重複序列發生了倒轉,使IRa與IRb變為同向排列[14]。反向重複序列可能可幫助維持葉綠體DNA的穩定[16]。
陸生植物新葉的葉綠體中一般有約100個葉綠體DNA,老葉中則僅剩15至20個[17],多包裹成擬核,一個葉綠體中常有數個擬核[10]。葉綠體DNA雖不與組蛋白結合[18],但紅藻的葉綠體DNA可編碼和組蛋白相似的組蛋白樣葉綠體蛋白(HC)與自身結合[19]。較原始的紅藻Cyanidioschyzon merolae(屬溫泉紅藻綱)的葉綠體擬核集中在葉綠體基質的中央,綠藻與陆生植物的葉綠體擬核則均勻分布於基質中[19]。
參見
编辑參考文獻
编辑- ^ Lyttleton, J. W. Isolation of ribosomes from spinach chloroplasts. Exp. Cell Res. 1962, 26 (1): 312–317. PMID 14467684. doi:10.1016/0014-4827(62)90183-0.
- ^ Heber, U. Protein synthesis in chloroplasts during photosynthesis. Nature. 1962, 195 (1): 91–92. Bibcode:1962Natur.195...91H. PMID 13905812. S2CID 4265095. doi:10.1038/195091a0.
- ^ Stocking, C. R.; Gifford, E. M. Incorporation of thymidine into chloroplasts of Spirogyra. Biochemical and Biophysical Research Communications. 1959, 1 (3): 159–164. doi:10.1016/0006-291X(59)90010-5.
- ^ Ris, H.; Plaut, W. Ultrastructure of DNA-containing areas in the chloroplast of Chlamydomonas. J. Cell Biol. 1962, 13 (3): 383–91. PMC 2106071 . PMID 14492436. doi:10.1083/jcb.13.3.383.
- ^ Shinozaki, K.; Ohme, M.; Tanaka, M.; Wakasugi, T.; Hayashida, N.; Matsubayashi, T.; Zaita, N.; Chunwongse, J.; Obokata, J.; Yamaguchi-Shinozaki, K.; Ohto, C. The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. The EMBO Journal. 1986, 5 (9): 2043–2049. ISSN 0261-4189. PMC 1167080 . PMID 16453699. doi:10.1002/j.1460-2075.1986.tb04464.x.
- ^ Ohyama, Kanji; Fukuzawa, Hideya; Kohchi, Takayuki; Shirai, Hiromasa; Sano, Tohru; Sano, Satoshi; Umesono, Kazuhiko; Shiki, Yasuhiko; Takeuchi, Masayuki; Chang, Zhen; Aota, Shin-ichi. Chloroplast gene organization deduced from complete sequence of liverwort Marchantia polymorpha chloroplast DNA. Nature. 1986, 322 (6079): 572–574 [2022-01-17]. Bibcode:1986Natur.322..572O. ISSN 1476-4687. S2CID 4311952. doi:10.1038/322572a0. (原始内容存档于2022-08-02).
- ^ Dann, Leighton. Bioscience—Explained (PDF). Green DNA: BIOSCIENCE EXPLAINED. 2002 [2022-01-17]. (原始内容存档 (PDF)于2012-09-17).
- ^ Clegg MT, Gaut BS, Learn GH, Morton BR. Rates and patterns of chloroplast DNA evolution. Proceedings of the National Academy of Sciences of the United States of America. 1994, 91 (15): 6795–801. Bibcode:1994PNAS...91.6795C. PMC 44285 . PMID 8041699. doi:10.1073/pnas.91.15.6795 .
- ^ 9.0 9.1 9.2 Shaw J, Lickey EB, Schilling EE, Small RL. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. American Journal of Botany. March 2007, 94 (3): 275–88. PMID 21636401. doi:10.3732/ajb.94.3.275.
- ^ 10.0 10.1 Burgess, Jeremy. An introduction to plant cell development. Cambridge: Cambridge university press. 1989: 62. ISBN 978-0-521-31611-8.
- ^ Daniell, H.; Lin, C.; Yu, M.; Chang, W. Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol. 2016, 17 (1): 134. PMC 4918201 . PMID 27339192. doi:10.1186/s13059-016-1004-2.
- ^ Clegg, M. T.; Gaut, B. S.; Learn, G. H.; Morton, B. R. Rates and patterns of chloroplast DNA evolution. PNAS. 1994, 91 (15): 6795–6801. Bibcode:1994PNAS...91.6795C. PMC 44285 . PMID 8041699. doi:10.1073/pnas.91.15.6795 .
- ^ 13.0 13.1 Berry, J. O.; Yerramsetty, P.; Zielinski, A. M.; Mure, C. M. Photosynthetic gene expression in higher plants. Photosynth. Res. 2013, 117 (1): 91–120. PMID 23839301. S2CID 16536768. doi:10.1007/s11120-013-9880-8.
- ^ 14.0 14.1 14.2 14.3 14.4 Sandelius, Anna Stina. The Chloroplast: Interactions with the Environment. Springer. 2009: 18. ISBN 978-3-540-68696-5.
- ^ 15.0 15.1 15.2 Kolodner R, Tewari KK. Inverted repeats in chloroplast DNA from higher plants. Proceedings of the National Academy of Sciences of the United States of America. January 1979, 76 (1): 41–5. Bibcode:1979PNAS...76...41K. PMC 382872 . PMID 16592612. doi:10.1073/pnas.76.1.41 .
- ^ 16.0 16.1 Palmer JD, Thompson WF. Chloroplast DNA rearrangements are more frequent when a large inverted repeat sequence is lost. Cell. 1982, 29 (2): 537–50. PMID 6288261. S2CID 11571695. doi:10.1016/0092-8674(82)90170-2.
- ^ Plant Biochemistry 3rd. Academic Press. 2005: 517. ISBN 9780120883912.
number of copies of ctDNA per chloroplast.
- ^ Biology 8th Edition Campbell & Reece. Benjamin Cummings (Pearson). 2009: 516.
- ^ 19.0 19.1 Kobayashi T, Takahara M, Miyagishima SY, Kuroiwa H, Sasaki N, Ohta N, Matsuzaki M, Kuroiwa T. Detection and localization of a chloroplast-encoded HU-like protein that organizes chloroplast nucleoids. The Plant Cell. July 2002, 14 (7): 1579–89. PMC 150708 . PMID 12119376. doi:10.1105/tpc.002717.